…um, at least as far as medical records go. SQL remains useful for a lot of other things, of course. But as far as electronic medical records are concerned, SQL is a really bad fit and should be taken out back and shot.
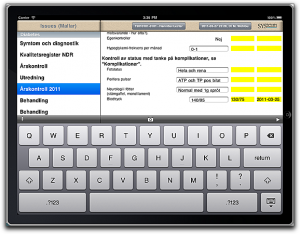
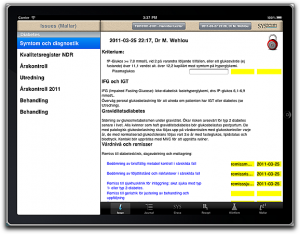
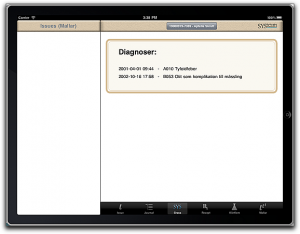
Medical records, in real life, consist of a pretty unpredictable stack of document types, so some form of graph database is very obviously the best fit for storage. Anything with rows and columns and predeclared types, is a very poor fit, except maybe for the patient demographics and lab data. Or maybe not even that.
The problem so far was the lack of viable implementations so instead of doing the right thing and creating the right database mechanism, most of us (me included) forced our data into some relational database, often sprinkling loose documents around the server for all those things that wouldn’t fit even if hit with a sledgehammer. All this caused mayhem with the data, concurrency, integrity, and not least, security.
I have to add here that I never personally committed the crime of writing files outside of the SQL, but squeezed them into the database however much effort it cost, but from the horrors I’ve encountered in the field, it seems not many were as driven as I was. I have, though, used Mickeysoft’s “structured storage” files for that, a bizarre experience at best.
You have to admit this is ridiculous. It leads to crazy bad databases, bad performance, horrible upgrading scenarios, and, adding insult to injury, high costs. Object-relational frameworks don’t help much and without going into specifics, I can claim they’re all junk, one way or the other, simply because the idea itself sucks.
From now on, though, there’s no excuse for cramming and mutilating medical records data into SQL databases anymore. Check out RDF first, to get a feel for what it can do. It’s part of the “semantic web” thing, so there’s a buzzword for you already.
A very good place to start is the rdf:about site, and right in the first section, there’s a paragraph that a lot of people involved in the development of medical records really should pause and contemplate, so let me quote:
What is meant by “semantic” in the Semantic Web is not that computers are going to understand the meaning of anything, but that the logical pieces of meaning can be mechanically manipulated by a machine to useful ends.
Once you really grok this, you realize that any attempts to make the computer understand the actual contents of the semantic web is meaningless. Not only is it far too difficult to ever achieve, but there is actually nothing to be gained. There’s nothing useful a computer can do with the information except present it to a human reader in the right form at the right time. What is important, however, is to let the computer understand just enough of the document types and links to be able to sort and arrange documents and data in such a way that the human user can benefit from accurate, complete, and manageable information displays.
It is almost trivial to see that this applies just as well to medical records. In other words, standardizing the terms of the actual contents of medical documents is a fool’s errand. It’s a pure waste of energy and time.
If a minimal effort would be expended to standardize link types and terms instead, we could fairly easily create semantic medical records, allowing human operators to utilize the available information effectively. All it would take for the medical community to realize this would be to raise their gaze and check out what the computer science community is doing with the web and then copy that. At least, that’s what we are aiming to do with the iotaMed project and I hope we won’t remain alone. What is being done using RDF on the web makes a trainload of sense, and we’re going to exploit that.
In practice, this means that you need to express medical data as RDF triples and graphs. This turns out to be nearly trivial, just as it is very easy to do for the semantic web. It’s a lot harder, and largely useless, for typical accounting data, flight booking systems, and others of that kind, but those systems should really keep using SQL as their main storage technique.
We also need a graph database implementation and we’re currently looking into Neo4j, an excellent little mixed license product that seems to fill most, if not all, requirements. But if it turns out it doesn’t, there are others out there, too. After all the years I’ve spent swearing at SQL Server and inventing workarounds for the bad fit, Neo4j and RDF is a breath of fresh air. The future is upon us, and it’s time to leave the computing middle ages behind us as far as electronic medical records are concerned.